Rice University’s breakthrough nanoporous silicon oxide technology for resistive random-access memory (RRAM) appears poised for commercialization.
Nanotechnology-based next generation memory nears mass production
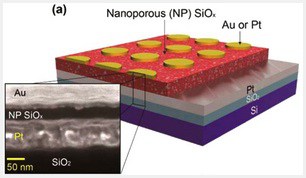

Rice University’s breakthrough nanoporous silicon oxide technology for resistive random-access memory (RRAM) appears poised for commercialization.
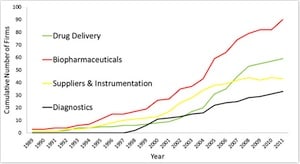
Study shows more than 500 firms involved in nanobiotechnology, which is expected to soon triple in size. Research points to the importance of broad networks and deep collaborations.

With biotech fundamental to several paths to advanced nanotechnology, a way to do biotech experiments in the cloud offers small groups the chance to quickly test their ideas.
B.R.AI.N.S., Berkeley BioLabs, and Foresight Institute to build an open source biological parts repository and design and distribute a line of “How-to Build Biological Machines” educational kits.

Foresight friends can use this discount to attend the SENS Rejuvenation Biotechnology Conference August 21-23, 2014 Santa Clara, California.

The photos from the 2014 Foresight Technical Conference highlight entrepreneurial efforts in space, biotechnology, and life extension.

A “sense of energy, momentum, and collegiality throughout the weekend” united attendees hearing about the integration of nano-engineered devices and materials into more complex systems, and the integration of nanoscale technologies into diverse applications.

At the 2013 Conference Philip Moriarty presented non-contact Atomic Force Microscope experiments demonstrating mechanical toggling of silicon dimers on a silicon surface. The crucial role of precise control of probe tip structure was emphasized.

Eric Drexler’s TEDx talk entitled “A Future of Radical Abundance: Transforming the Material Basis of Civilization” is available for viewing on Youtube as well as on Drexler’s blog site. As described by the Oxford Martin School, where Drexler is a scholar with the Programme on the Impacts of Future Technology: Dr. Eric Drexler’s talk from… Continue reading TEDx talk: "Transforming the Material Basis of Civilization"
An invitational workshop to address the opportunities and challenges of nanotechnology for developing countries will be held in parallel with Foresight’s open nanotechnology conference “The Integration Conference”.