A commentary over at Gizmodo argues that ideas about molecular manufacturing that sounded like science fiction in 1986 now sound more like science fact.
Atomically precise manufacturing as the future of nanotechnology

A commentary over at Gizmodo argues that ideas about molecular manufacturing that sounded like science fiction in 1986 now sound more like science fact.
The idea that nanorobots fabricated by atomically precise manufacturing processes are a likely part of our future, and that this is a good thing, is appearing more frequently, largely as a result of Drexler’s recent book Radical Abundance.
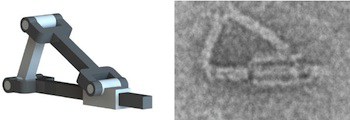

Scaffolded DNA origami is combined with hinges of single- or double-stranded DNA to built simple machines parts that have been combined to program simple to complex motions.
One example is presented of how well the meme is spreading that nanotechnology will evolve toward atomically precise manufacturing that will in turn bring forth a world of abundance.
Combinations of different types of DNA nanorobots, implementing different logic gates, work together to tag a specific type of cell in a living cockroach depending on the presence or absence of two protein signals.
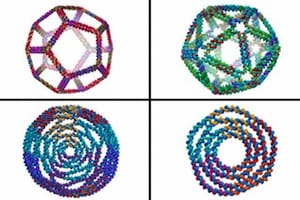
A more general computational framework predicts the structures of 2D and 3D-curved DNA nanostructures impossible to predict using previously available computational methods. May lead to 3D-printing DNA nanostructures?
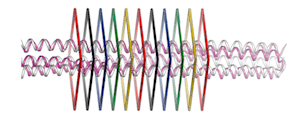

Advances in the de novo design of coiled-coil proteins made by two different research groups proceeding by two different routes demonstrate that the range of protein nanostructures potentially available for various molecular machine systems is significantly larger than the range of such structures already exploited by natural selection.
The US National Science Foundation announced a new grant program to develop and apply next-generation networking to advance nanotechnology and other emerging technologies to meet important national needs.

A small, interactive group of invited experts gathered in Palo Alto recently to discuss prospects for revolutionary advances in energy storage, transmission, and generation through nanotechnology.

The need for improved imaging and characterization on the nanoscale was emphasized in the 2007 Roadmap and again at the 2013 Foresight Conference on Atomic Precision. We noted last year a new advancement in atomic-scale resolution of 10-nm platinum particles, requiring multiple imaging techniques in combination, and recently the marked improvement in optical imaging for… Continue reading A Breakthrough in 3D Imaging by EM Alone