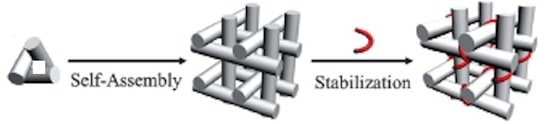
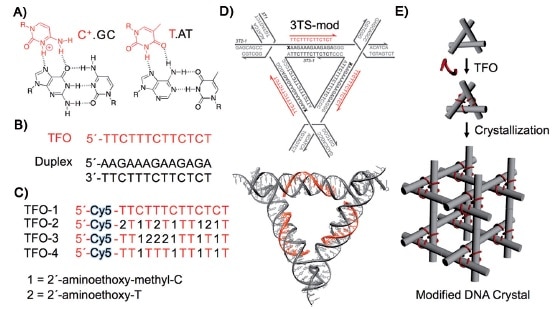
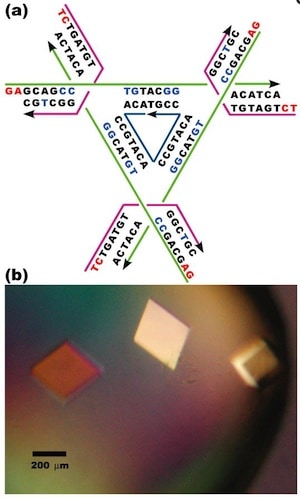
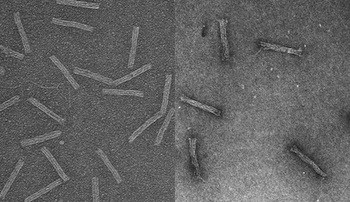
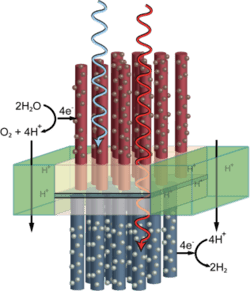
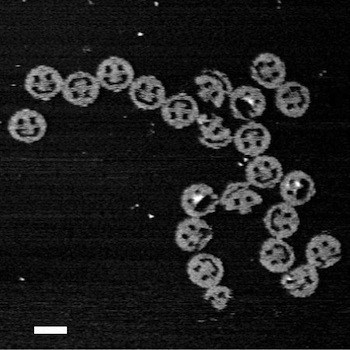
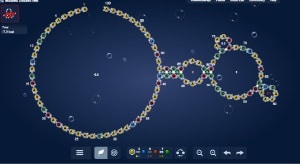
A DNA strand capable of forming a triple helix with a portion of the DNA double helices in a macroscopic DNA crystal enhances the weak interactions holding the crystal together so that the crystal remains stable in the absence of a high ionic strength environment.
Triple helices stabilize macroscopic crystals for DNA nanotechnology