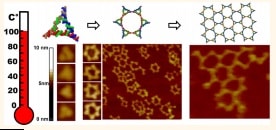
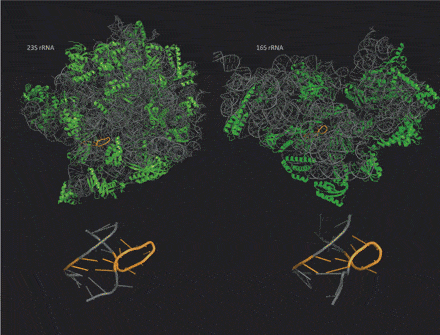
The complex molecular recognition code of RNA offers RNA nanotechnology a greater variety of 3D structures and functions than are present in DNA nanotechnology, but the RNA structures can be fragile. New RNA triangles that resist boiling solve this problem.
Robust triangular RNA brick adds to RNA nanotechnology toolkit